MedEQ Tutorial#
Here is a minimal, but complete example showing the main interface to the medeq.MED object.
import medeq
import numpy as np
# Create DataFrame of MED free parameters and their bounds
parameters = medeq.create_parameters(
["velocity", "viscosity", "radius"],
minimums = [-9, -9, -9],
maximums = [10, 10, 10],
)
def instrument(x):
'''Example unknown "experimental response" - a complex non-convex function.
'''
return x[0] * np.sin(0.5 * x[1]) + np.cos(1.1 * x[2])
# Create MED object, keeping track of free parameters, samples and results
med = medeq.MED(parameters)
# Initial parameter sampling
med.sample(24)
med.evaluate(instrument)
# New sampling, targeting most uncertain parameter regions
med.sample(16)
med.evaluate(instrument)
# Add previous / manually-evaluated responses
med.augment([[0, 0, 0]], [1])
# Save all results to disk - you can load them on another machine
med.save("med_results")
# Discover underlying equation; tell MED what operators it may use
med.discover(
binary_operators = ["+", "-", "*", "/"],
unary_operators = ["cos"],
)
# Plot interactive 2D slices of responses and uncertainties
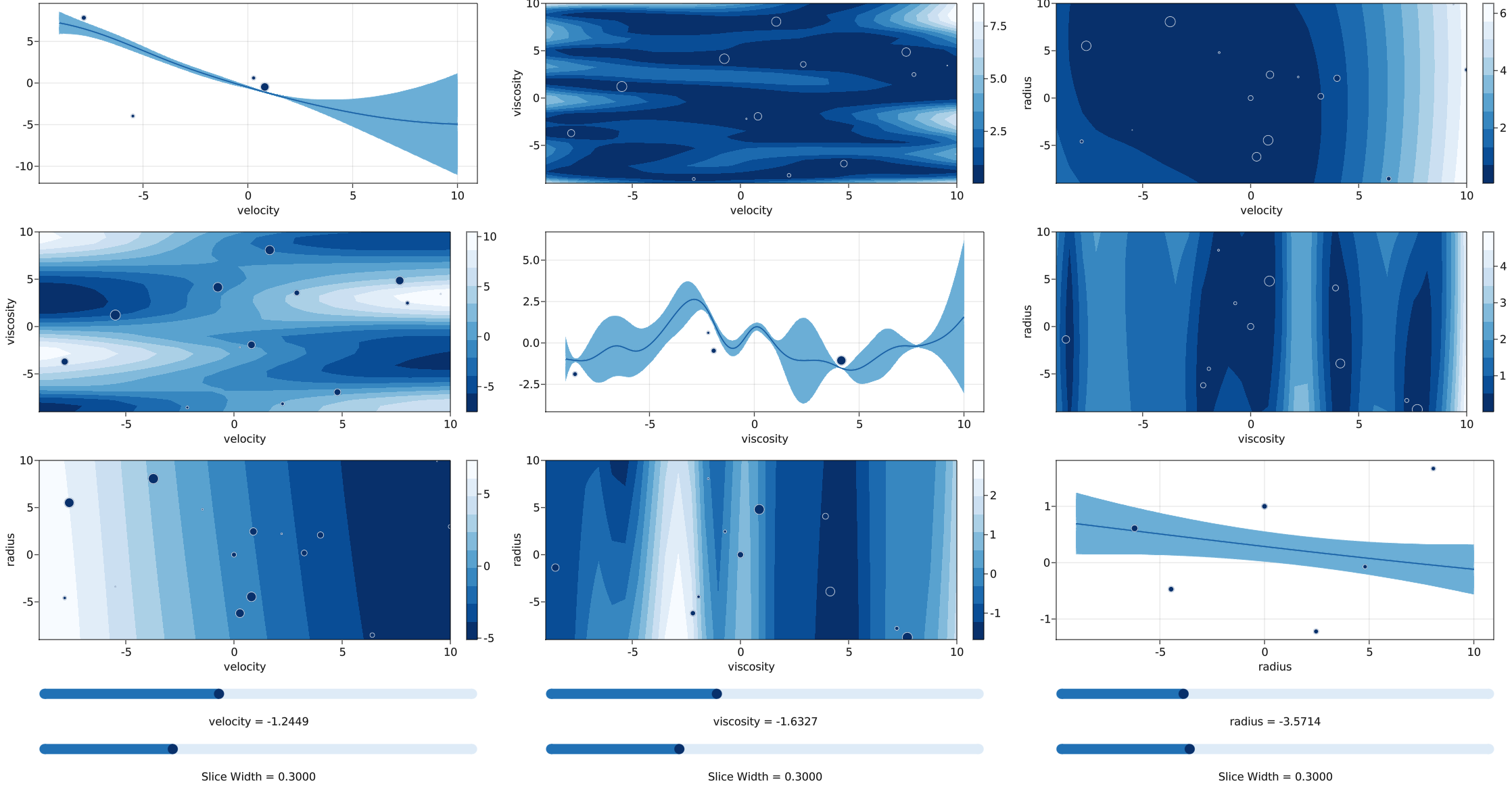
med.plot_gp()
Here are the equations found by SymbolicRegression.jl at various complexity levels:
==============================
Hall of Fame:
-----------------------------------------
Complexity Loss Score Equation
1 1.656e+01 1.025e-07 -0.19632196
2 1.626e+01 1.812e-02 cos(radius)
3 1.541e+01 5.332e-02 (-0.20152433 * velocity)
4 1.227e+01 2.278e-01 (velocity * cos(velocity))
6 8.668e+00 1.739e-01 (velocity * cos(-1.0899653 * velocity))
8 4.988e-01 1.428e+00 (velocity * cos(1.5935777 + (-0.50125474 * viscosity)))
10 4.946e-01 4.271e-03 ((-1.016915 * velocity) * cos(7.8330894 + (0.5005289 * viscosity)))
11 1.241e-01 1.383e+00 (cos(radius) + (velocity * cos(1.5515859 + (-0.49880704 * viscosity))))
13 0.000e+00 1.151e+01 (cos(1.1000026 * radius) + (velocity * cos(1.5707898 + (-0.50000036 * viscosity))))
Note how it discovered the sin(x) term as cos(1.57 + x).